D-amino acids in protein synthesis
Can tRNA select the L-enantiomer?
As a general rule, D-AAs do not bind tRNA molecules; thus, their presence in proteins is related to posttranslational modifications (Doolitle, 1983).
Nonetheless, it was recently demonstrated that the cellular translational machinery accepts D-AA-tRNAs into the ribosomal aminoacyl-tRNA binding site A, uses it as a peptidyl-transfer acceptor, and translocates the resulting peptidyl-D-AA-tRNA to the ribosomal P (peptidyl) site. Subsequently, the latter traps the ribosomal peptidyl-transferase center in a conformation that impairs peptidyl transfer (Englander et al., 2015)
Chiral proofreading during translation of the genetic code
Proteins are essentially homochiral in nature since they are composed of only L-amino acids (besides glycine, which is achiral). However, the cellular pool consists of significant amounts of various free D-amino acids (D-AAs) that do have important physiological roles; for instance, D-serine and D-aspartate act as neurotransmitters in neurons (Dunlop et al., 1986; Hashimoto et al., 1992). It is necessary to exclude D-AAs from the translational machinery because heterochiral polypeptides cannot take on a proper 3D form due to geometric constraints and therefore undergo spontaneous misfolding and degradation. Multiple steps during translation act as cellular checkpoints to prevent D-AAs from infiltrating proteins. Nevertheless, some of the aminoacyl-tRNA synthetases (aaRSs) – which carry out the first step of translation by coupling amino acids to their cognate tRNAs – are not adequately enantioselective and thus sometimes mischarge D-AAs on tRNAs. An enzyme, named D-tyrosyl RNA deacylase (DTD), which could remove D-tyrosine and D-phenylalanine mischarged on tRNATyr and tRNAPhe, respectively, was discovered in 1967 by Calendar and Berg (Calendar and Berg, 1967). Later biochemical characterization showed that DTD can act on multiple D-aminoacyl-tRNAs (Soutourina et al., 1999), and the enzyme was then renamed D-aminoacyl-tRNA deacylase (the abbreviation DTD was, however, retained). Hence, DTD was implicated in perpetuating homochirality in proteins, and its activity on D-aminoacyl-tRNAs was referred to as “chiral proofreading” (Ahmad et al., 2013). Structural studies deciphered the mechanistic basis underlying DTD enantioselectivity, wherein an invariant cross-subunit Gly-cisPro motif was found to be responsible for the same (Fig. 1) (Ahmad et al., 2013). Further investigations demonstrated that chiral proofreading site for DTD uses only L-chiral rejection and not D-chiral selection as the design principle to ensure substrate specificity (Fig. 1).
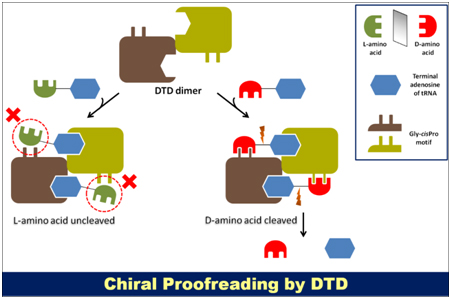
Fig. 1. Mechanistic basis of DTD’s enantioselectivity. DTD uses an invariant cross-subunit Gly-cisPro motif to ensure substrate specificity through L-chiral rejection.
This naturally led to the finding that DTD has significant activity on the achiral cognate substrate Gly-tRNAGly, thereby engendering the “glycine misediting” paradox. It was found that elongation factor thermo unstable (EF-Tu), which delivers aminoacyl-tRNAs to ribosome, protects the achiral substrate against DTD and therefore resolves the paradox (Fig. 2). However, the concentration of DTD in the cell must be tightly regulated because higher DTD levels can overcome the protection given to Gly-tRNAGly by EF-Tu, thereby causing cellular toxicity (Fig. 2) (Routh et al., 2016). Another interesting aspect about DTD is that the side chains in the active site are dispensable; DTD uses main-chain atoms for substrate selectivity and the 2′-OH of 3′-terminal A76 of tRNA for catalysis (Routh et al., 2016). DTD is thus a primordial RNA-based catalyst in which the protein acts as a scaffold to provide an environment conducive for the aminoacyl-tRNA to perform (substrate-assisted) catalysis.
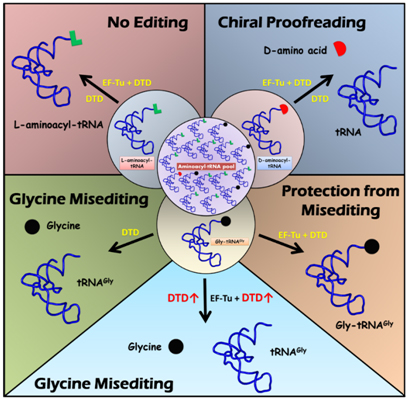
Fig. 2. Model for the “glycine misediting” paradox and its resolution. DTD efficiently decouples D-aminoacyl-tRNAs, but does not act on the L-counterparts. In the absence of EF-Tu, DTD deacylates (hydrolyzes) Gly-tRNAGly, thereby generating the “glycine misediting” paradox. EF-Tu is able to resolve this problem by protecting the achiral substrate. However, if DTD levels become high, it is able to relieve this protection, thus causing cellular toxicity.
References
Ahmad S, Routh SB, Kamarthapu V, Chalissery J, Muthukumar S, Hussain T, Kruparani SP, Deshmukh MV, Sankaranarayanan R., Mechanism of chiral proofreading during translation of the genetic code. eLife. 2013; 2:01519. doi: 10.7554/eLife.01519.
Calendar R, Berg P. D-Tyrosyl RNA: formation, hydrolysis and utilization for protein synthesis. J Mol Biol. 1967; 26(1):39–54. doi: 10.1016/0022-2836(67)90259-8.
Doolittle, R. (1983) Probability and the origin of life. In: Godfrey, L.R., ed., 1983. Scientists Confront Creationism, W.W. Norton, NY
Dunlop DS, Neidle A, McHale D, Dunlop DM, Lajtha A., The presence of free D-aspartic acid in rodents and man. Biochem. Biophys. Res. Commun. 1986; 141(1):27–32. doi: 10.1016/S0006-291X(86)80329-1.
Hashimoto A, Nishikawa T, Hayashi T, Fujii N, Harada K, Oka T, Takahashi K., The presence of D-serine in rat brain. FEBS Lett. 1992; 296(1):33–6. doi: 10.1016/0014-5793(92)80397-Y.
Routh SB, Pawar KI, Ahmad S, Singh S, Suma K, Kumar M, Kuncha SK, Yadav K, Kruparani SP, Sankaranarayanan R., Elongation factor Tu prevents misediting of Gly-tRNA(Gly) caused by the design behind the chiral proofreading site of D-aminoacyl-tRNA deacylase. PLoS Biol. 2016; 14(5):e1002465. doi: 10.1371/journal.pbio.1002465.
Soutourina J, Plateau P, Delort F, Peirotes A, Blanquet S., Functional characterization of the D-Tyr-tRNATyr deacylase from Escherichia coli. J. Biol. Chem. 1999; 274(27):19109–14. doi: 10.1074/jbc.274.27.19109.
Authors
Rajan Sankaranarayanan, CSIR-CCMB – Hyderabad
Loredano Pollegioni, Università degli Studi dell’Insubria
